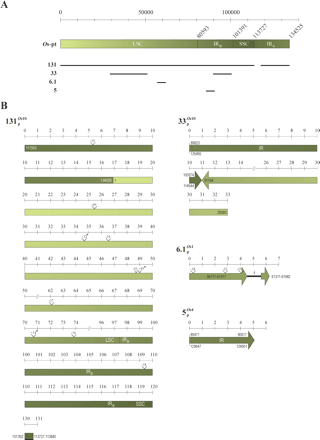
Structure of long continuous insertions of ptDNA in the nuclear genome of O. sativa. (A) Structure of rice ptDNA (Os-pt). The position of the long (LSC) and short (SSC) single-copy regions, as well as of the two inverted repeats (IRA and IRB) are indicated. The four nuclear insertions (see bottom) are depicted as black lines below their regions of origin in the rice plastid chromosome. When the origin of a fragment
of nuclear ptDNA was ambiguous, i.e., when derived from repetitive regions, it was assigned to the repeat located most 5′
of the organellar DNA. For instance, NUPTs solely homologous to IR-specific sequences were always assigned to IRB.(B) Structure of the four nuclear insertions of rice ptDNA 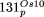 ,
, 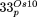 ,
, 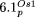 and
and 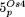 . Coloring and numbers indicate the position of the homologous sequence in the plastid chromosome based on the structure reported
in A. The orientation of NUPTs relative to ptDNA is indicated by arrowheads; arrowheads pointing to the left indicate a reverse orientation. In the case of the IR regions of the ptDNA, the position of both homologous ptDNA sequences
is provided. Fragments of ptDNA deleted from the nuclear integration are indicated by triangles with the size of the deletion
indicated (e.g. “–14”) (see Table 3); short insertions or duplications are indicated accordingly (e.g. “+45”; duplications are highlighted by an asterisk; Supplemental
Table 2). For
. Coloring and numbers indicate the position of the homologous sequence in the plastid chromosome based on the structure reported
in A. The orientation of NUPTs relative to ptDNA is indicated by arrowheads; arrowheads pointing to the left indicate a reverse orientation. In the case of the IR regions of the ptDNA, the position of both homologous ptDNA sequences
is provided. Fragments of ptDNA deleted from the nuclear integration are indicated by triangles with the size of the deletion
indicated (e.g. “–14”) (see Table 3); short insertions or duplications are indicated accordingly (e.g. “+45”; duplications are highlighted by an asterisk; Supplemental
Table 2). For 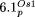 , the long insertion of nonorganelle DNA is indicated by a black line and the letter “i” (Supplemental Table 1).
, the long insertion of nonorganelle DNA is indicated by a black line and the letter “i” (Supplemental Table 1).











