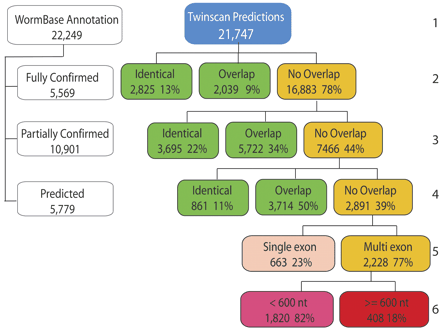
Breakdown of genome-wide predictions by TWINSCAN 2.01 in comparison to the WS130 annotations. (Row 1) Total number of WormBase annotations and TWINSCAN predictions. (Row 2) Breakdown of TWINSCAN predictions into those that are identical to fully confirmed WormBase predictions, those that overlap but are not identical, and those that do not overlap (orange). (Row 3) Breakdown of TWINSCAN predictions that do not overlap fully cDNA-confirmed ORFs by comparison to the partially cDNA confirmed WormBase ORFs. (Row 4) Breakdown of TWINSCAN predictions that do not overlap fully or partially confirmed WormBase ORFs by comparison to predicted WormBase ORFs. (Row 5) Breakdown of TWINSCAN predictions that do not overlap any of the above into single exon (beige) and multiexon (orange) predictions. (Row 6) Breakdown of novel multiexon TWINSCAN predictions into those that are shorter than 200 amino acids (pink) and those that are at least 200 amino acids (red). Analysis of predictions by an earlier and slightly less accurate version of TWINSCAN (2.0α), by comparison to WS100 ORFs, placed 265 novel ORFs of at least 200 amino acids in the red box, all of which were tested experimentally.











