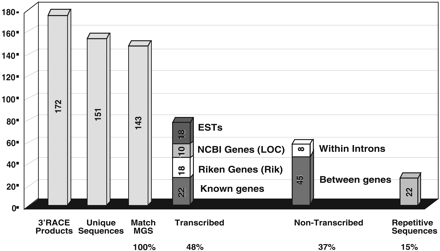
Tagged-sequences mutagenesis with the LNPAT1 3′ gene trap. Here 172 vector-fusion transcripts cloned by 3′-RACE corresponded to genes disrupted in ES cells by the LNPAT1 vector as determined by their DNA sequence (confirmed 3′-RACE products). Of these, 151 provided unique sequence tags (i.e., they were not derived from sister clones from the same culture dish), and 143 matched the mouse genome sequence (MGS). Based on the mouse genome sequence annotation, 68 (48%) of the RACE sequences matched transcribed sequences (i.e., exons), corresponding to 22 previously characterized genes, 18 Riken cDNAs, 10 transcription units identified by gene prediction software (NCBI LOC genes), and 18 EST contigs not included among the Entrez gene annotations (ESTs). Of the remaining 53 sequence tags, eight matched intron sequences within annotated genes, 45 matched genomic sequences not contained within annotated transcription units and therefore may be between genes, and 22 contained repetitive sequences and could not be placed on the genome sequence.











