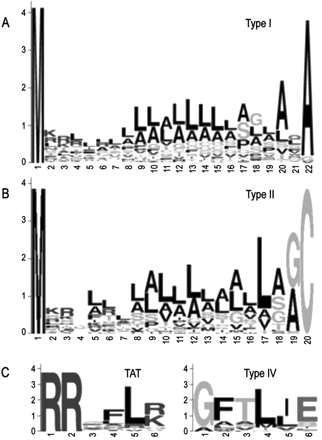
Amino acid sequence logos of predicted P. aeruginosa PAO1 type I signal peptides (A), type II signal peptides (B), as well as the TAT and type IV motifs (C). For type I, all signal sequences where the cleavage site predictions were in agreement and that were 22 amino acids in length (mode) were used (n = 77). For type II, all lipoproteins 19 amino acids in length (mode) plus the Cys residue were used (n = 31). All predicted TAT motifs (n = 27) and type IV motifs (n = 23) were used.











