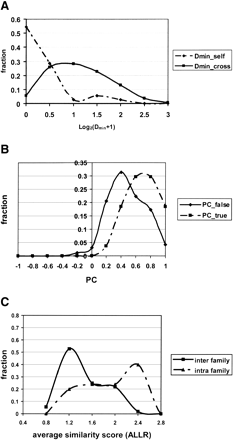
Three types of information. (A) Two types of normalized D distributions. Dmin_self: the distance between a TF gene and its closest binding site in the genome. Dmin_cross: the distance between a TF gene and the closest site for a different TF. (B) Two types of phylogenetic correlation (PC) distributions. PC_true: PC for true TF-DNA-motif pairs. PC_false: PC for false TF-DNA-motif pairs. (C) Distribution of average similarity scores for motifs from the same family and from different families. Motifs were aligned using an ungapped Smith-Waterman algorithm and scored using the ALLR statistic.











