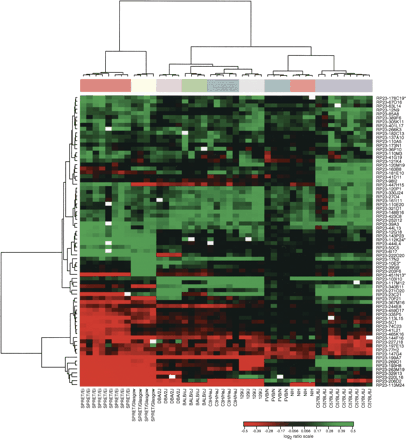
Agglomerative hierarchical clustering of 74 regions of sequence variation. We used a Euclidian metric and Ward method to cluster both samples and clones. For overlapping BACs in contigs, we used the one clone from the contig for which the greatest number of observations met the threshold (clone denoted by an asterisk). Note that all mice cluster in two separate branches according to species in agreement with accepted phylogeny. Within the M. spretus cluster, all SPRET/Ei mice cluster together and are more closely related to one another than to the outbred SPRET/Glasgow mice. In the M. musculus branch, all individual mice from each of the strains cluster together. Furthermore, FVB/N and NIH are closest to each other in the clustering dendrogram, which again agrees with the strain phylogeny.











