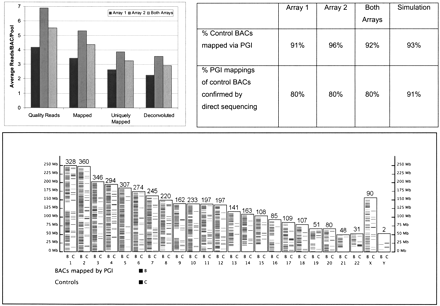
Mapping rhesus BACs onto human genome. On average, 5.5 quality reads were obtained per BAC per pool, 4.3 (78%) could be mapped, 3.2 (58%) uniquely mapped, and 2.9 (53%) deconvoluted (top left). Because each BAC was pooled four times, in two row pools and two column pools using the transversal design, the read numbers need to be multiplied by four to obtain number of reads per BAC. A total of 3858 BACs mapped onto 4178 loci (bottom; an interactive version of this figure is available at www.genboree.org). Number of mappings appears above each chromosome. Performance of PGI was evaluated by using 99 randomly selected BACs and by simulation experiments (top right), as described in the Methods section. Each of the 99 BACs was directly sequenced and mapped. A PGI mapping of a BAC was counted as confirmed if the BAC was mapped to the same position by directly obtained sequence.











