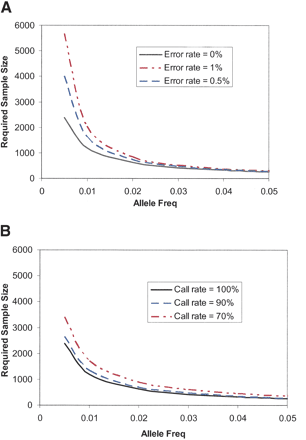
The effect of inaccurate genotypes (A) and incomplete genotyping (B) on the number of patients required to have 80% power to find a genetic association. A genetic model has been assumed in which the relative risk of the causative allele (GRR) is two. The effect is assumed to be multiplicative. The causative allele frequency is plotted on the x-axis. The largest loss of power comes with making inaccurate calls for markers with low frequency. By contrast, incomplete data result in smaller loss of power, which is felt across the allele frequency spectrum. The data from the MIP assay are accurate enough to be used for the investigation of rare alleles without significant loss of power.











