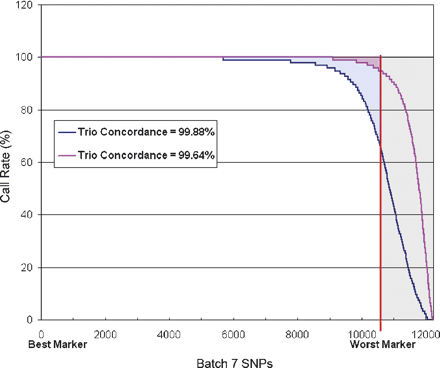
The effect of clustering parameters on performance metrics. In this plot, the markers for Batch 6 are ordered along the x-axis such that the marker with the highest call rate is at the origin, while the worst performing of the ∼12,000 markers is at the right. The y-axis shows the call rate for each of these markers across 95 individuals. The markers that exhibit poor call rates are called nonconverted and are shown in the gray area. The red curve shows a choice of cluster calling parameters that emphasizes high completeness by accepting calls on the periphery of clusters. More markers show very high call rates and the amount of missing data shown by the red shaded region is minimal (99.2% completeness). The overall accuracy as measured by trio concordance shows that a small number of erroneous calls are being made (99.64% concordance). If one wishes to eliminate these incorrect calls, the base caller can be tuned to be more stringent. This choice allows very high accuracy (∼99.9% trio concordance) while causing more missing data (blue shaded region). The choice of cluster calling parameters should thus be chosen according to the intended use of the data.











