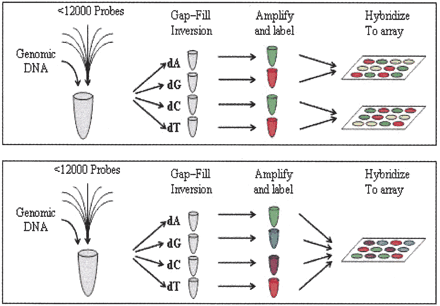
Schematic of the MIP assay process. MIP reactions are set up adding an enzyme mix and genomic DNA to the probe pool. This mix is then split into four tubes, each receiving a distinct nucleotide species. After gap-filling and probe inversion, inverted probes are amplified using common PCR primers. These amplicons are labeled using one of two labeling processes. In the two-color labeling scheme (top), the A and C reactions are labeled with one fluorophore while the G and T reactions are labeled with a spectrally distinct fluorophore. The A and G reactions are then pooled and hybridized to one tag array while the C and T reactions pooled and hybridized to a second array. Both arrays are then scanned using a GeneChip array scanner in two spectral channels to generate four fluorescent signals for each tag. In the four-color labeling scheme (bottom), each of the four reactions is labeled with a spectrally distinct fluorophore. All four reactions are then pooled and hybridized to a single tag array which is scanned using a GeneChip AT CCD imager in four spectral bands. In both cases four images are generated containing the four allele signals for each SNP marker.











