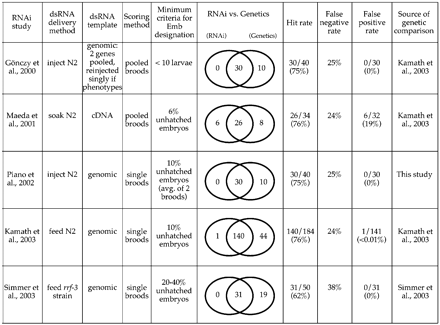
Summary of large-scale RNAi studies. First five columns: For each published RNAi study, the dsRNA delivery method, the template used for generation of dsRNA, the scoring method, and the minimum criteria for scoring are indicated (for reviews of large-scale RNAi projects, see Piano and Gunsalus 2002 and Sugimoto 2004). The scoring method refers to whether individual broods or composites of two or more broods were scored to obtain the percentage of unhatched embryos. The scoring criteria indicate the minimum level of lethality that must be observed in order to score a gene as embryonic lethal. For the Gönczy study, three injected hermaphrodites were placed on a single plate and allowed to lay eggs for 48 h; plates with fewer than 10 hatched larvae were scored as embryonic lethal. Because over 100 eggs are commonly produced by three animals during this time period, in this study the cutoff for scoring lethality was ∼90% unhatched embryos. The Simmer study used the rrf-3 strain, which shows some embryonic lethality and sterility in the absence of RNAi; thus their cutoff for scoring lethality was more stringent than most of the other studies. Last five columns: Comparison of RNAi phenotypes with phenotypes from genetic mutations for genes from each RNAi study that are represented by characterized mutant alleles. Venn diagrams indicate genes identified as Ste or Emb by RNAi only (left; potential false positives), by both RNAi and genetics (overlap), or by genetic analysis only (right; potential false negatives). We define the “hit rate” as the number of Ste or Emb genes identified by RNAi relative to the number of genes with known sterile or lethal genetic alleles. We define the “false positive rate” as the number of genes with Ste or Emb phenotypes by RNAi that do not give rise to sterility or lethality according to genetic analysis, divided by the total number of RNAi-lethal genes represented by genetic alleles.











