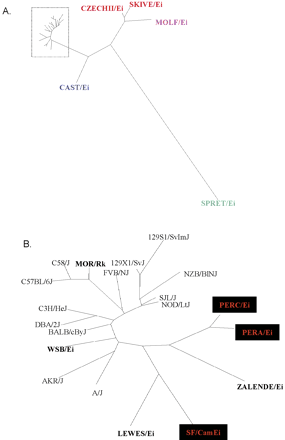
Phylogenetic tree constructed with 1004 SNPs discovered from resequencing of 13 laboratory inbred strains and 12 wild-derived inbred strains (labeled in bold) from five subspecies: musculus (SKIVE/Ei and CZECHII/Ei), molossinus (MOLF/Ei), castaneus (CAST/Ei), M. spretus (SPRET/Ei), and domesticus (LEWES/Ei, MOR/Rk, WSB/Ei, ZALENDE/Ei, PERA/Pk, PERC/Ei, and SF/CamEi). The phylogeny was constructed using the dnadist program in the PHYLIP package (http://evolution.genetics.washington.edu/phylip.html). The branch length information was used to plot evolutionary distance between strains in the DrawTree program. (A) Phylogeny of all strains. (B) Detailed phylogenetic tree of species in the dotted rectangle in A.











