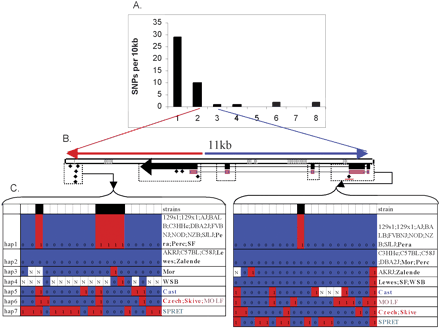
(A) SNP density (defined as the number of SNPs per 10 kb) of a 70-kb haplotype block. The x-axis labels the 10-kb intervals in this block. (B) The 11-kb genomic region within this block encodes mRNA NM_145481. The region was split into two sections labeled with red and blue lines to represent SNP-rich and SNP-poor segments, respectively. (Gray boxes) Regions with repetitive sequence. The mRNA is transcribed in reverse orientation, and the exons are labeled with black rectangles. (Magenta) The coding regions. (Black diamonds) The SNPs in the Celera data. The last SNP in this region (with a red underline) is the only missense variation in this region. (Dotted rectangles) The regions selected for resequencing. (C) Haplotypes constructed with SNPs discovered by resequencing. (Black) The SNPs discovered in the five Celera strains. The two alleles of a SNP are labeled as 0 or 1; (N) an ambiguous allele. The wild-derived inbred strains are labeled in bold, with colors indicating strains of the domesticus (black), musculus (red), molossinus (magenta), and castaneus (blue) subspecies. (Green) M. spretus species.











