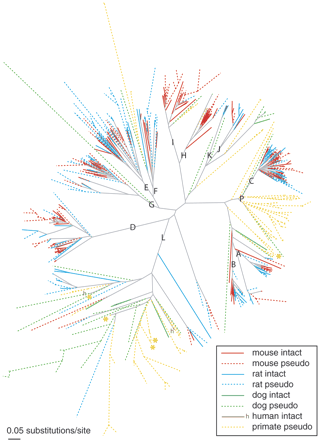
Neighbor-joining gene tree illustrating subfamily clades (A-L, P) of intact and pseudogene V1R-like sequences identified in the genomes of five mammalian species. Mouse intact (red), mouse pseudogene (red dashed), rat intact (blue), rat pseudogene (blue dashed), dog intact (green), dog pseudogene (green dashed), human intact (brown, denoted “h”), and human/chimpanzee pseudogene (yellow dashed) branches are shown. Two chimpanzee and three human pseudogenes that were previously annotated as intact V1Rs (see Methods) are indicated with yellow and orange asterisks, respectively. A total of 87 mouse, 39 rat, 27 dog, 59 chimpanzee, and 58 human V1R-like pseudogenes were excluded from this tree, because the length of confidently aligned sequence was too short (see Methods). A list of these excluded sequences, as well as their presumed subfamily (based on closest fast×34 matches), is provided in Supplemental Figure D.











