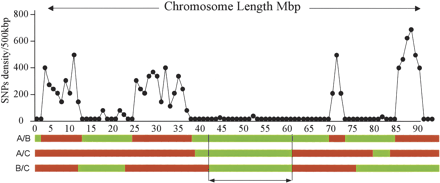
The SNP density between any two inbred strains of mice varies according to the chromosomal region concerned and changes abruptly when passing from one region to the next. This observation is in agreement with historical data on the origin of laboratory inbred strains indicating that they are derived from a small pool of wild ancestors belonging to different subspecies of the genus Mus. Because of this polyphyletic origin, the mouse genome can be regarded as a mosaic of chromosomal segments of various sizes. When the SNP density is low, the segments in question share the same ancestral origin and the few observed SNPs are those resulting from recent mutations. When the SNP density is high, on the contrary, the chromosomal segments have a different origin. When three strains are compared, as on the diagram represented here, one can perfectly observe that three homologous segments have a high SNP density on pairwise comparisons, if all three of them have an independent origin stemming, for example, in three different subspecies of the genus Mus. When a particular region with low SNP density cosegregates with a particular phenotype, the region in question may harbor the genetic determinants for the phenotype in question. (Redrawn with permission from PNAS © 2003, Wiltshire et al. 2003.)











