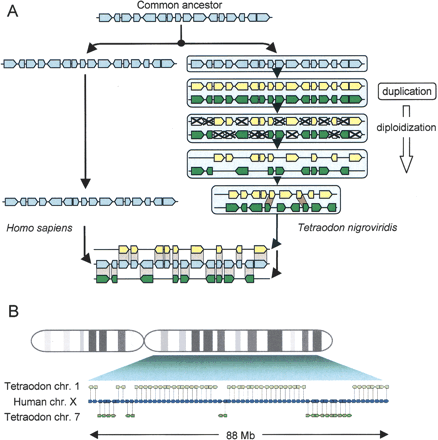
(A) Duplication leads to double-conserved synteny. After speciation, a chromosome or chromosome segment from an ancestral species is duplicated in one lineage but not in the other. In the former (e.g., Tetraodon), supernumerary copies of genes are progressively deleted from each of the duplicated segments in approximately equal proportion (diploidization). Ultimately the two duplicated chromosomes only contain 50% of the initial gene complement and are thus very different from each other. The difficulty of finding the original pair of sister chromosomes can be alleviated by a comparison with a genome that originates from the same pre-duplication ancestor, but that did not duplicate (e.g., human). The nonduplicated chromosome segment should contain genes with orthologs alternating between the two duplicated chromosomes (adapted from Kellis et al. 2004). (B) Example of double-conserved synteny. An 88-Mb region covering the majority of the long arm of human chromosome X contains 65 genes with orthologs in the Tetraodon genome, alternating between chromosome 1 and chromosome 7. Genes are represented by small arrows that indicate the orientation of transcription.











