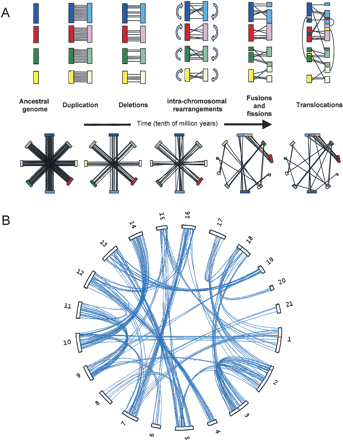
(A) A schematic model of whole-genome duplication with four chromosomes, followed by massive gene loss, chromosome fusions and fissions, inter- and intrachromosomal rearrangements. Each colored rectangle is a chromosome, and lines are drawn between duplicate copies of genes present on sister chromosomes or chromosome segments. The top panel starts with the sister chromosomes facing each other and illustrates the changes induced by chromosome rearrangements, while the lower panel shows the situation in a circular representation, which assumes one does not know the relationships between chromosomes a priori. After several million years of evolution, the distribution of the few duplicate genes that remain do not bear a trace of the ancient duplication event. (B) The same representation as in the lower panel in (A), but with real data from the Tetraodon genome: Despite more than 300 million years since the duplication, the distribution of about 2% of Tetraodon genes that remain strictly in two copies (joined by blue lines) in the genome shows a striking pattern where chromosomes are associated in pairs (e.g., chromosome 9 and 11, or 10 and 14), or sometimes in triplets (e.g., chromosomes 5, 13, and 19). The former suggests that no interchromosomal rearrangements have occurred on these chromosome pairs since the duplication, while the latter is reminiscent of a chromosome fusion or fission.











