Unraveling genomic regulatory networks in the simple chordate, Ciona intestinalis
Abstract
The draft genome of the primitive chordate, Ciona intestinalis, was published three years ago. Since then, significant progress has been made in utilizing Ciona's genomic and morphological simplicity to better understand conserved chordate developmental processes. Extensive annotation and sequencing of staged EST libraries make the Ciona genome one of the best annotated among those that are publicly available. The formation of the Ciona tadpole depends on simple, well-defined cellular lineages, and it is possible to trace the lineages of key chordate tissues such as the notochord and neural tube to the fertilized egg. Electroporation methods permit the targeted expression of regulatory genes and signaling molecules in defined cell lineages, as well as the rapid identification of regulatory DNAs underlying cell-specific gene expression. The recent sequencing of a second Ciona genome (C. savignyi) permits the use of simple alignment algorithms for the identification of conserved noncoding sequences, including microRNA genes and enhancers. Detailed expression profiles are now available for almost every gene that encodes a regulatory protein or cell-signaling molecule. The combination of gene-expression profiles, comparative genome analysis, and gene-disruption assays should permit the determination of high-resolution genomic regulatory networks underlying the specification of basic chordate tissues such as the heart, blood, notochord, and neural tube.
The ancestral chordate gave rise to three groups, i.e., the vertebrates, cephalochordates, and tunicates (which include the ascidians). While the vertebrates diversified into many familiar aquatic and terrestrial species, the cephalochordates and tunicates remained in the ocean and evolved into highly specialized filter feeders. In the tunicates, acquisition of an endoglucanase gene and cellulose tunic (possibly derived by lateral gene transfer from bacteria) (see Dehal et al. 2002; Matthysse et al. 2004; Nakashima et al. 2004) led to the evolution of a highly divergent adult body plan lacking almost all coelomic cavities. However, the morphogenesis of chordate structures (e.g., notochord and dorsal nerve cord) in tunicate tadpoles has been well conserved despite the “retrograde” evolution of adult anatomy.
The recent sequencing of two closely related ascidian species, Ciona intestinalis and Ciona savignyi, has revived the status of the ascidians as a major developmental model system (Dehal et al. 2002; Satoh 2003; Satoh et al. 2003). Research has confirmed that many aspects of ascidian and vertebrate embryogenesis rely on a conserved set of orthologous genes and cellular processes (Satoh 2003; Passamaneck and Di Gregorio 2005). This is clearly the case for development of the nervous system (Meinertzhagen et al. 2004), notochord (Passamaneck and Di Gregorio 2005), and heart (Satou et al. 2004) and may also be true for other tissues and organs, including the blood cells, pharyngeal gill slits, endostyle, and neural crest (Jeffery et al. 2004). However, ascidian embryos are extraordinarily simple, with low cell numbers, rapid development, and well-defined lineages (Fig. 1A,B) (Satou et al. 2004). The compact genomes of both Ciona species also facilitate the identification of regulatory DNAs (e.g., enhancers) that direct tissue-specific and lineage-specific patterns of gene expression (Johnson et al. 2004; Kusakabe 2005). In addition, genetic simplicity of the Ciona genomes results from a general restriction in gene duplication events (Dehal et al. 2002). For example, there are approximately four vertebrate homologs of the heart homeobox gene Tinman, whereas Ciona possesses only one such gene (Wada et al. 2003). Thus, the paucity of redundant paralogs for key patterning genes (encoding regulatory proteins or cell-signaling molecules) simplifies the assessment of their function. Additionally, the ability to manipulate isolated blastomeres and to create transgenic embryos through electroporation of fertilized eggs greatly increases the potential to conduct in-depth analyses of Ciona development. Thus, research in Ciona presents an extraordinarily powerful avenue for deciphering fundamental chordate developmental networks.
There have been a number of recent reviews assessing different aspects of ascidian research in the post-genomic era (Canestro et al. 2003; Satoh 2003; Satoh et al. 2003; Du Pasquier 2004; Kasahara et al. 2004; Meinertzhagen et al. 2004; Kusakabe 2005; Passamaneck and Di Gregorio 2005). In order to avoid redundancy, this review is focused primarily on how the latest research can be compiled in the form of genomic regulatory networks. We also describe research on the cell biological output of these networks. We have structured the review around discussion of three studies that serve as entry points into the state of Ciona research following the sequencing of the genome. These studies span three distinct approaches (and continents). The first study is representative of a “systems approach” to unravel complex developmental processes, specifically the early specification of various endomesodermal lineages (Imai et al. 2004). The second study exemplifies the ability of Ciona researchers to make rapid progress in dissecting complex regulatory networks underlying the development of conserved chordate tissues, such as the dorsal nerve cord (Bertrand et al. 2003). The third study illustrates the forward genetics approach and how exploitation of Ciona's morphological simplicity can provide deep insights into genetic regulation of cellular processes (Jiang et al. 2005). Along with a detailed overview of these studies, each section will also use these studies as launching points to discuss other related work and the advantages and challenges inherent to each of these approaches.
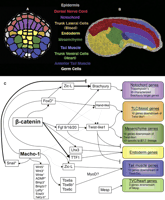
Network map for specification of endomesodermal lineages. (A) Diagram of the vegetal hemisphere of a Ciona gastrula. Each cell is color coded according to its lineage as indicated by the color labels in the center. (B) Confocal image of an early tailbud embryo stained with phalloidin. Embryonic tissues have been artificially colored in accordance with the color scheme in A to display the fate of each of the lineages. Domains of some tissues are approximate and may not be wholly accurate. (C) Summary of data on interactions between genes involved in specification of endomesodermal lineages. Boxes for each lineage are colored according to the scheme in A and B. Bold lines indicate that this interaction has been verified through the manipulation of binding sites within an enhancer. Asterisks indicate evidence for appropriate binding sites in the noncoding DNA upstream of the target genes. Dotted lines indicate tangential interaction indicated by embryological manipulations. The notochord determinant Brachyury encodes a T-box homeodomain protein. The genes required for specifying different mesodermal lineages (NoTric [previously known as Handlike], Twist-like1, Mesp, MyoD) all encode bHLH transcriptional factors. Further details regarding endomesodermal differentiation can be found in Supplement 1.
Comprehensive studies of gene expression and function
Since the sequencing of the genome, the potential of Ciona as a model system has been greatly enhanced by a number of systematic annotation studies (Imai 2003; Satou and Satoh 2003; Yagi et al. 2004). Of particular importance was the recent publication of gene-expression patterns for nearly every identified regulatory gene and signaling factor in the C. intestinalis genome (Imai et al. 2004). This survey focused mostly on early stages from the zygote through gastrulation, but also included representative stages (neurula, early and late tailbud) from older embryos. This work supplements the sequencing of an extensive set of tissue and stage-specific EST libraries and expression surveys of 1043 transcripts from these libraries (Satou et al. 2002b) along with thorough annotation of predicted genes (Satou and Satoh 2003). Together, these data constitute one of the most comprehensive sets of expression patterns for any model organism and have enormous utility for studies in Ciona and other chordates.
An overview of this new expression data generated two important observations regarding early Ciona development (Imai et al. 2004). First, over 70% of the genes encoding transcription factors and signaling molecules are maternally expressed. The substantial presence of maternally loaded factors may underlie rapid determination of cell fates in early Ciona embryos. Second, only 65 transcription factors and 25 signaling factors are zygotically expressed by the early gastrula stage. As almost all embryonic tissues are specified by this time, it is suggested that “comprehensive transcriptional networks” can be constructed based on the interactions between these 90 genes. Based on this prediction, a quantitative real time PCR-based assay was conducted, measuring the response of these 90 genes to suppression of the characterized tissue-specification factors β-catenin, FoxD, and FGF9/16/20 (Imai et al. 2004).
The comprehensive expression screen and associated RT-PCR assays complement an extensive body of research focused on early specification of Ciona endomesoderm lineages. We have attempted to draw together much of this work into a summary network (Fig. 1C). On the right are the regulatory outputs that subdivide the vegetal hemisphere of the 110-cell Ciona embryo into endoderm and five distinct mesodermal lineages. Details of the genes that control the formation of these lineages and the regulatory interactions can be found in Supplement 1. The subdivision of distinct endomesoderm lineages depends on two crucial maternal factors, Macho-1 and β-catenin (Fig. 1C). β-catenin is translocated into the nuclei of vegetal blastomeres, where it plays an essential and apparently highly conserved role in endomesoderm specification (Imai et al. 2000). Macho-1 functions as the maternal determinant for the tail-muscle lineage, a role that probably evolved within the tunicates in association with the extremely rapid embryogenesis of a functional tail (Nishida and Sawada 2001; Sawada et al. 2005).
To date, the majority of research on these early networks has involved loss-of-function studies to identify potential targets. There are relatively few studies in which direct interactions between regulatory genes and their targets have been verified by the manipulation of specific binding sites within defined enhancers (Fig. 1C, shown by dark lines). Characterization of cell-specific enhancers for the core patterning genes, ZicL, FoxD, FGF9/16/20, Tbx6, Twist-like1, Mesp, MyoD, and NoTric, would provide the basis for establishing causal interconnections among these genes. A recent study on the Ciona Otx enhancer demonstrates the potential of such an approach (Bertrand et al. 2003).
Regulatory interactions underlying Ciona neural specification
Despite the simplicity of the Ciona larval central nervous system (∼330 cells), the overall organization is comparable to the vertebrate CNS (Fig. 2B) (Meinertzhagen et al. 2004). Many aspects of early neural specification and later neuronal differentiation appear to rely on conserved chordate gene networks that have been expanded during the evolution of more complex vertebrate neural structures (Meinertzhagen et al. 2004). Thus, dissection of gene networks in Ciona neurogenesis will provide crucial insights into conserved aspects of these networks in vertebrates. This potential is exemplified by a recent study on the regulation of the early neural specification gene Otx (Bertrand et al. 2003), which encodes a homeodomain protein expressed in both Ciona and vertebrate anterior neural structures.
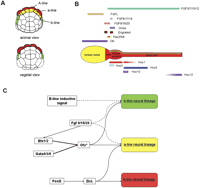
Regulatory network for Ciona neural development. (A) Animal and vegetal views of 64-cell stage embryo showing the position of A (red), a (yellow), and b-line (green) neural lineages. (B) The Ciona CNS is subdivided into sensory vesicle, neck, visceral ganglion, and nerve cord along the A-P axis. The progeny of three neural lineages are colored according to the scheme in A. Five Hox genes exhibit partial spatial colinearity along the visceral ganglion and nerve cord (Ikuta et al. 2004). Additional homeodomain transcriptional factors like Otx, Pax2/5/8, and Dmbx are expressed in the anterior CNS (Takahashi and Holland 2004). Four FGF genes are expressed in partially overlapping domains along the CNS (Imai et al. 2002; Satou et al. 2002a). (C) Regulatory network for neural specification. The three Ciona neural lineages are specified by a combination of maternal and zygotic factors. Bold lines indicate that this interaction has been verified through the manipulation of binding sites within an enhancer. Asterisks indicate evidence for appropriate binding sites in the noncoding DNA upstream of the target genes. Dotted lines indicate tangential interaction indicated by embryological manipulations. Further details of the differentiation the neural lineages can be found in Supplement 2.
The methodology of this study represents an optimal use of Ciona's potential to dissect cis-regulation of gene networks. A 3.5-kb enhancer was identified for the Ci-Otx gene, and subsequent manipulations characterized an ∼120-bp activation element (a-element). This analysis exploited two key features of the Ciona system as follows: (1) the Ciona genome is very compact, with regulatory regions often located immediately upstream of the genes they control; (2) in Ciona, regulatory regions can be rapidly assessed and dissected in great detail by high-throughput electroporation of reporter constructs into fertilized eggs (Corbo et al. 1997; Harafuji et al. 2002; see Fig. 3A,B). The authors utilized the 120-bp Otx regulatory element to identify the endogenous activator as Ci-FGF9/16/20. Based on previous work, it was clear that Otx induction relied on MEK signaling (Hudson and Lemaire 2001; Hudson et al. 2003). However, identification of the endogenous factor relied on two further advantages of Ciona. (1) The limited number of paralogs in the Ciona genome reduces the number of candidate factors (six Ciona FGFs vs. 22 vertebrate FGFs). (2) The low cell number of early Ciona embryos permits stringent evaluation of candidate factors through in situ hybridization. (The subsequent publication of the comprehensive gene expression database discussed above greatly magnifies this advantage, see Fig. 4).
At this point, the authors successfully addressed a pivotal obstacle in unraveling signaling networks, i.e., how cells of various lineages differentially interpret a broad signal. In Ciona, FGF9/16/20 is known to mediate induction of the three following lineages: notochord, mesenchyme, and neural cells (Imai et al. 2002; Hudson et al. 2003; see Figs. 1,2). To reveal the mechanistic basis for a neural-specific response to this signal, the authors used phylogenetic footprinting. The availability of both C. intestinalis and C. savignyi genome assemblies provides an opportunity to identify potential regulatory DNAs by simply aligning orthologous sequences from the two species (Johnson et al. 2004). Alternatively, the high level of polymorphism between individuals from different populations of C. intestinalis can also be used to identify regulatory sequences (Boffelli et al. 2004). The authors used the cross-species comparison to reveal a concentration of potential transcriptional factor binding motifs for Ets and Gata factors. They were then able to rapidly confirm the importance of these sites through mutational analysis and by creating versions of the a-element containing only Ets or Gata sites. This analysis showed that the FGF signal causes general activation of Otx, while the requirement for Gata coactivation limits this response to the emerging neural lineage. Once again, the ability to perform such in-depth analysis relied on the ability to test reporter constructs by high-throughput electroporation (Fig. 3C). We wish to emphasize that reporter gene activity is monitored in the “zero” generation, just hours after electroporation of 1-cell embryos as compared with the lengthy process of creating germ-line transformation in vertebrate systems. Mosaic expression is not a problem, due to the efficient incorporation of the fusion genes and the small number of cell divisions leading to the formation of the tadpole. Additionally, it is now possible to establish stable transgenic lines through injection or electroporation of recombinant genes flanked by the Minos transposon (Sasakura et al. 2003; Matsuoka et al. 2005). This technique has been used for the identification of regulatory DNAs via enhancer trapping (Awazu et al. 2004).
The results discussed above provide a definitive regulatory network for activation of Otx in the anterior neural lineage (the a-line lineage). The posterior CNS in Ciona is derived from two distinct lineages, the A-line and b-line (Fig. 2A,B). The regulatory networks specifying these two neural lineages are less well understood and may involve cell-autonomous mechanisms and cell-cell interactions, respectively (for this and other additional details concerning Ciona neural development, see Supplement 2). The comprehensive analysis of Otx regulation demonstrates the power of the Ciona system for the detailed dissection of cis-regulatory elements. The challenge is to extend this methodology to help decipher other specification networks underlying early development. Such approaches are now being applied to the early development of the heart (B. Davidson, W. Shi, and M. Levine, in prep.), endoderm (Satou et al. 2001; Fanelli et al. 2003; Imai 2003; Oda-Ishii et al. 2005), tail muscles (Yagi et al. 2005), and stomodaeum (Christiaen et al. 2005). In combination with the comprehensive gene-expression studies and functional assays discussed previously, these enhancer studies will set the stage for deciphering the basic gene networks in the Ciona embryo. However, such networks are only the first step in understanding Ciona development. The next step is to investigate the roles of down-stream genes in influencing the cellular components underlying tissue morphogenesis. In other words, how do interconnected networks of regulatory factors and cell-signaling molecules control the detailed behavior of individual cells? As discussed in the next section, Ciona is also an excellent model system for exploring these downstream cellular processes.
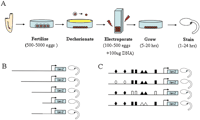
Using electroporation to transiently express plasmid DNA in Ciona embryo. The Ciona system allows rapid delivery and expression of reporter gene constructs in the developing embryo. It is also convenient to use tissue-specific enhancers for targeted misexpression in specific embryonic lineages. (A) Eggs and sperm are isolated from the adult for in vitro fertilization. The fertilized eggs are dechorionated, mixed with plasmid DNA and electroporated at the 1-cell stage. The electroporated embryos are then allowed to develop to appropriate developmental stages for reporter gene analysis (X-gal staining) or fixed for in situ hybridization. (B) Schematic of serial deletion analysis. The last construct depicts the minimal enhancer element (solid line) activating reporter gene expression with a basal promoter. It often requires testing of 20-40 deletion constructs to identify a minimal enhancer. (C) Schematic of site-directed mutagenesis of a minimal regulatory element. Shapes represent different putative transcription factor binding sites, filled for wild-type and unfilled for mutated. In this case, the binding sites represented by diamonds were required for reporter expression. Thorough characterization of a minimal enhancer through this method would also require testing of at least 20-40 constructs. Because one can test 10-16 constructs in a single round of electroporations, a single researcher can test over 100 constructs/week. Thus, in-depth serial deletion (B) and subsequent site-directed mutagenesis (C) can be conducted over a few weeks rather than many months.
Notochord morphogenesis
During Ciona notochord development, many of the genes downstream of Brachyury appear to control cellular processes such as adhesion, intercalation, and cell shape (Hotta et al. 2000). Studies on notochord differentiation and morphogenesis highlight the broad range of tools that are available for the dissection of complex morphogenetic processes (Di Gregorio et al. 2002; Keys et al. 2002). A recent study used forward genetics (Fig. 5) combined with detailed analysis of mutant cell behavior to characterize the role of Prickle, a component of the planar polarity pathway, in the morphogenesis of the notochord (Jiang et al. 2005).
The authors began with a screen for spontaneous mutations, focusing on defects in notochord formation. Because the larval body is largely disposable, Ciona embryos with severe defects in larval structures (including the notochord and dorsal nerve cord) can still metamorphose into reproductive juveniles. The authors isolated the aimless (aim) mutant based on severe defects in larval tail formation. A variety of genomic tools identified aim as a mutation in the conserved cell polarity gene, Prickle. This gene was previously identified in a comprehensive subtractive hybridization screen for notochord-specific genes downstream of Brachyury (Takahashi et al. 1999).
To explore the precise role of Prickle in notochord formation, the authors compared the behavior of isolated notochord precursor cells from mutant and wild-type embryos. Early lineage determination in Ciona embryos permits the isolation of individual blastomeres from defined lineages. Confocal analysis led to the observation that mutant cells exhibit loss of localized bipolar protrusions. The transparency and low cell number of Ciona embryos make it feasible to observe cell behavior in vivo. The authors exploited high-throughput Ciona transgenesis to target tagged transcripts of Prickle and the conserved cell polarity gene Dishevelled to the developing notochord. Careful observation of wild-type and mutant transgenic embryos provided evidence that Prickle mediates the localization of Dishevelled along the medio-lateral axis of intercalating notochord cells.
The power of Ciona for determining the genetic basis of cellular processes is only now being exploited. There is great potential in applying similar techniques to other aspects of Ciona development, including directed cell migration of heart and blood cells, cell movements underlying gastrulation, and morphogenesis of the neural tube.
Emerging resources and techniques
There are a number of significant resources and techniques that will promote further research. Some of these tools are currently being developed, while others are still under consideration.
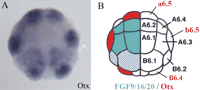
Use of in situ expression patterns to identify candidate genes. (A) In 32-cell stage Ciona embryos, Otx is expressed in the neural lineage a6.5 and b6.5 blastomeres (photo taken from the Ghost database). (B) Diagram of a 32-cell stage embryo illustrating that at this stage FGF9/16/20 is the only Ciona FGF zygotically expressed in cells (blue) neighboring Otx expressing cells (red) (Bertrand el al. 2003). Multiplex in situ hybridization techniques (B. Davidson, W. Shi, and M. Levine, in prep.) that permit visualization of multiple genes simultaneously will greatly enhance the ability to identify potential regulatory cascades based on comparative expression patterns.
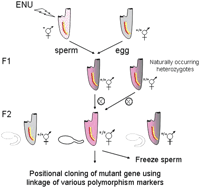
Forward genetic approach to identify recessive mutations in Ciona. Ciona mutagenesis involves a classical F2 screen (Moody et al. 1999). The F0 animals are treated with the chemical mutagen ENU and maintained for several weeks to reduce mosaicism. The mutagenized sperm is then crossed to wild-type animals to obtain heterozygous F1 progenies. The F1 animals are individually self-fertilized to produce F2 progeny that are screened for phenotypic abnormalities. The sperm of mutant lines can then be cryo-preserved for long-term storage. Alternatively, naturally occurring recessive mutations can be isolated by self-crossing single animals from the wild population. The point mutation can be determined by various mapping methods, including linkage analysis of amplified length polymorphism (Jiang et al. 2005).
Web resources
The Ciona intestinalis genome assembly and annotation is provided by the Joint Genome Institute at http://genome.jgi-psf.org/ciona4/ciona4.home.html. The newly released assembly version 2.0 contains 141 M base pairs of non-N sequences (A. Sidow, pers. comm.) and is a significant improvement over the original version 1.0, which covers only about 2/3 of the predicated genome (Dehal el al. 2002). The Ghost database (http://ghost.zool.kyoto-u.ac.jp/indexr1.html) provides the transcriptional factor/signaling molecule expression profiles and the cDNA and EST database information. In addition to these two most widely accessed databases, a number of computer resources are under development.
-
An integrated and searchable database of in situ expression profiles is currently under development (http://aniseed-ibdm.univ-mrs.fr). This database will include detailed three-dimensional representations of expression patterns using precisely staged embryos. This will be particularly useful, as there is currently no systematic, consistent staging criteria for Ciona embryos following the onset of gastrulation.
-
A searchable database of the Ciona savignyi genome was recently placed in Ensembl (http://www2.bioinformatics.tll.org.sg:8082/Ciona_savignyi/) and the Ciona intestinalis genome has been incorporated into the Vista browser (http://pipeline.lbl.gov/cgi-bin/gateway2). The Vista browser provides easy acquisition of orthologous Ciona savignyi sequences needed for the phylogenetic identification of regulatory DNAs.
-
A reference assembly of the Ciona savignyi genome was produced by joint efforts of the Broad Institute (Vinson el al. 2005) and Stanford University. The annotated version 2.0 covering ∼170 Mb will soon be available for download (A. Sidow, pers. comm.).
-
The current version of the genome did not incorporate the extensive EST database in its gene predictions. More accurate annotations of many genes are available on the Ghost database (see above). Efforts are also underway at Ensembl to annotate both new Ciona genome assemblies based on gene predication programs, the EST database, and the comparisons between the two Ciona genomes (A. Sidow, A. Ureta-Vidal, pers. comm.).
-
Efforts are underway to produce improved genome assembly and scaffolds for each of the 14 chromosomes (Shoguchi et al. 2004). Due to the high sequence polymorphism of the C. intestinalis genome, repeat sequences were not assembled in the initial assembly, resulting in shorter genomic scaffolds (Dehal et al. 2002). Comparisons to the C. savignyi genome should also greatly assist in a more complete assembly.
-
In Ciona mRNA 5′-leader, trans-splicing may be a common occurance (Vandenberghe et al. 2001) and thus may represent a challenge for accurate annotation of full-length transcripts. Efforts to systematically obtain full 5′ sequences for annotated genes are currently underway.
Other resources
-
DNA oligo microarrays covering the whole transcriptome are under development to replace the existing cDNA microarray, which has only about 85% coverage (Azumi et al. 2003). Although the gene expression survey discussed above (Imai et al. 2004) represents a powerful resource, it suffers from inherent problems associated with in situ hybridization, namely, that the quality of probes vary, leading to background noise that may obscure the true expression pattern. For example, in comparing the gene expression survey (Imai et al. 2004) with the study on Macho-1 targets (Yagi et al. 2004), it becomes clear that a large number of transcription/signaling factors expressed by the 32-cell stage (and subsequently down-regulated by Macho-1 suppression) were not detected by in situ hybridization until after the 110-cell stage. Extensive microarray screens using staged embryos and isolated blastomeres will provide a more complete assessment of the gene networks underlying embryogenesis.
-
Although the Forkhead promoter has been used to visualize the activities of a variety of tissue-specific enhancers, it is far from ideal, since it sometimes causes spurious expression patterns (Di Gregorio et al. 2001). The Brachyury promoter may be better suited for some enhancer analysis (Bertrand et al. 2003), but there is still a great need for additional core promoters.
-
Recently completed EST libraries of early juvenile stages should help launch studies into the differentiation of critical post-larval structures such as the endostyle, branchial gill slits, and heart. Many of the putative regulatory genes that are not expressed in the embryo might be essential for post-metamorphic events.
-
Establishment of ascidian stock centers will overcome the main obstacle to a robust Ciona research community, which is year-round access to gravid adults. Most research takes place in “land-locked” laboratories, and it is not always easy to find animals (particularly in winter months) that provide embryos for electroporation assays. To overcome this, centers where wild-type and stable transgenic lines are developed and maintained are under development in both Japan (Shimoda) andthe USA (UC Santa Barbara). Such centers will be critical for providing gravid animals year round and for distribution of stable transgenic lines. Particularly useful lines might include Gal4 UAS transgenics for expression of constructs in particular lineages and lines with GFP-tagged lineages for easy assessment of mutant phenotypes.
Conclusions
The Ciona system possesses virtually every modern analytical tool for the comprehensive determination of the genomic regulatory networks underlying development. We anticipate that some of the current deficiencies, such as access to gravid animals and the availability of inexpensive microarrays, will be remedied in the near future. It is easy to envision complete networks governing the specification, differentiation, and morophogenesis of key chordate tissues and organs such as the blood, heart, brain, nerve cord, and notochord. These networks should provide the foundation for unraveling the more elaborate genetic interactions used for the construction of homologous structures in vertebrates. In principle, it should be possible to integrate gene expression profiles, interlocking networks of regulatory genes and cell-signaling pathways to identify the on/off state of every gene in the Ciona genome in every blastomere at each stage in embryogenesis.
Footnotes
-
[Supplemental material is available online at www.genome.org.]
-
Article and publication are at http://www.genome.org/cgi/doi/10.1101/gr.3768905.
-
↵1 Corresponding author. E-mail bandl{at}calmail.berkeley.edu; fax (510) 643-5785.
- Cold Spring Harbor Laboratory Press