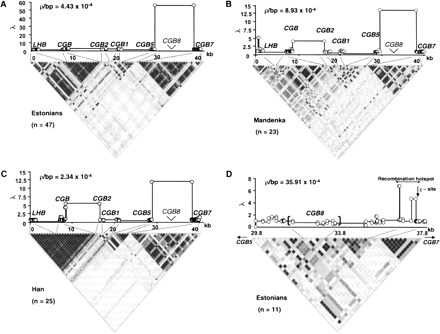
LD structure by r2-blot and estimation of crossing-over activity (measured by ρ =4Necbp) and potential recombination hotspots (measured by [λ = ρ between a locus pair]/[average ρ for the analyzed region]) by the PHASE algorithm (Stephens et al. 2001) across the LHB/CGB genome cluster for SNPs (○) with MAF > 10%: (A) Estonians; (B) Mandenka; and (C) Han Chinese samples. Regions of predicted high crossover activity colocalize with the breakdown (white) LD areas and low recombination rate with strong allelic associations (black/dark gray). (D) Refinement of recombination hotspot by resequencing genomic region between CGB5 and CGB7 genes in an Estonian sample as a model. The recombination hotspot colocalizes with LD breakdown and is located between CGB8 and CGB7 within a <1-kb region next to a crossover-associated Escherichia coli χ-sequence.











