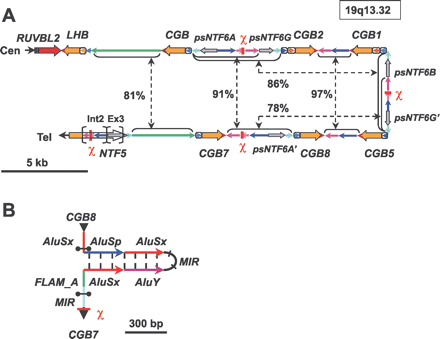
(A) Structure of the human LHB/CGB gene cluster. Identical color codes refer to highly homologous DNA sequences within the cluster. Genes (RUVBL2, LHB/CGBs, NTF5) are depicted as wide arrows in the direction of transcription with promoters as boxed arrows 5′ of the genes. Intergenic areas are marked as narrow lines, indicating also the direction of a segment, and broken arrows unite intergenic regions that share >75% of sequence identity. χ denotes the localization of consensus Escherichia coli χ-sequence (GCTGGTGG) (Smith 1988) associated with crossing-over and gene-conversion activity. Int2 and Ex3 refer to intron 2 and exon3 in the NTF5 gene; the former is the source of intergenic regions between CGB genes, and the latter has given rise to NTF6 pseudogenes. (B) Prediction of single-stranded DNA secondary structure for the estimated recombination hotspot (bordered by black brackets) between CGB8 and CGB7. The hotspot is flanked by double inverted Alu-sequences forming the stem (625 bp) of the palindrome, and its center falls within the loop (222 bp). (MIR) Mammalian-wide interspersed repeat; (FLAM_A) Alu-element-like repeat.











