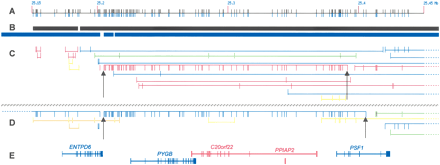
The 20p11.21 region of chromosome 20 that contains large 72 and 76 giSNP clusters in the UK unrelateds and Asian samples, respectively. From top to bottom: (A) SNP location of all SNPs genotyped and present in each population; (B) Gabriel et al. (2002) defined haplotype blocks for UK unrelateds (black) and Asians (blue); (C,D) giSNP clusters from UK unrelateds (above black-hashed line) and Asian (below black-hashed line) populations, respectively. GiSNPs in the same cluster are connected by a horizontal bar and MAF is represented by color (MAF: <10, blue; 10–20, green; 20–30, yellow; 30–40, orange; 40+, red). The horizontal bar between the giSNP ticks represents the number of SNPs per cluster. The higher the bar, the greater the number of giSNPs per cluster (bar height maximum, 15 giSNPs). The start and finish points of the 72- and 76-member giSNP clusters are indicated by black arrows; (E) gene locations.











