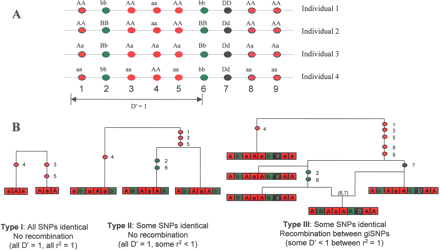
Schematic representations of the three different types of giSNP cluster. (A) Genotype representation of the categories for nine SNPs genotyped on four illustrative individuals. Category 1 giSNP clusters (red SNPs 3, 4, and 5) do not have any intervening SNPs that are nonindistinguishable. Category 2 clusters (green SNPs 2 and 6) have intervening SNPs, but there is no evidence of recombination throughout the cluster (i.e., all SNPs in the region are D′ = 1). Category 3 clusters (red SNPs 1, 3, 4, 5, 8, and 9) contain intervening SNPs that are nonindistinguishable, as well as evidence for a recombination event in the region (i.e., some pairs of SNPs are D′ < 1). (B) Bifurcating genealogical trees (Type I and Type II) and ancestral recombination graph (Type III) describing formation of the three types of giSNP cluster. Each graph describes the ancestral events giving rise to a giSNP cluster. Colored circles in the graph represent an ancestral mutation with the number indicating the SNP involved. The (6,7) in the graph of the Type III giSNP cluster refers to an ancestral recombination event between SNPs 6 and 7. The boxes at the base of the figure refer to the multilocus haplotype present in the population sample, with each box representing a single SNP with alleles coded as a capital vs. lower case letter for each cluster. Since genotypes are formed through the pairing of haplotypes, a pair of loci will be genetically indistinguishable when only two pair-wise conformations are present in the population (i.e., aa and AA are the only pair-wise combinations present among all haplotypes; or alternatively, aA and Aa are the only combinations present in the population). In the case of the Type I cluster, all SNPs are genetically indistinguishable. In the Type II cluster, all SNPs have D′ = 1 with each other. SNPs 2 and 6 are genetically indistinguishable, but are separated by the Type I subcluster consisting of SNPs 3, 4, and 5. In the Type III cluster, SNPs 1, 3, 4, 5, 8, and 9 are genetically indistinguishable, but are separated by SNP 7, which is different from any other SNP, and forms all four possible haplotypes with any other SNP in the cluster. Note that in this example the Type I cluster is part of a larger Type III cluster. These would not be listed as separate in this study and are shown separately in this figure for descriptive purposes only.











