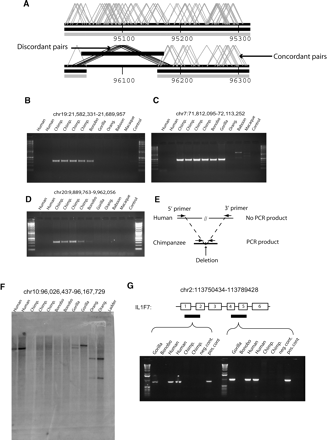
Detection and validation of “chimpanzee deletions.” (A) An example of a chimpanzee deletion event mapped to its corresponding position on human chromosome 10 (build34 coordinates in kb). Two criteria were used to identify chimpanzee deletions: multiple discordant (>49.5 kb) fosmid pairs (black angled lines covered by the black bar) and the absence of concordant fosmid pairs (gray lines) within the region. (B–D) Oligonucleotide sequences (Supplemental Table 5) were designed in regions of conserved human–chimpanzee sequence flanking each deletion breakpoint (see schematic in panel E). PCR products corresponding to the expected size were detected in chimpanzee but not human due to the increased distance between annealing oligonucleotides in the human genome. Results from other closely related apes and Old World monkeys provide outgroup information regarding lineage-specificity of the event. Bands of unexpected size are products of non-specific binding in more distant species. Panel C shows the deletion of a region on chromosome 7 that contains four human genes; POM121, WBSCR20C, TRIM50C, and FKBP6. (E) A schematic of the PCR primer design in chimpanzee and human. (F) Probes for Southern hybridization were developed based on human sequence corresponding to the predicted site of the deletion (see Methods; Supplemental Table 5) and hybridized against a primate panel of restriction-digested primate DNA. The probes successfully hybridized to human genomic DNA but not chimpanzee genomic DNA. Bands of different sizes and lighter intensity in more distant species likely show mutations in restriction enzyme sites. This panel shows a region that contains the human gene CYP2C18 on chromosome 10. (G) The results of an RT-PCR amplification of peripheral blood RNA from exons 1–2 and 3–4 in the IL1F7 gene on chromosome 2 in primates, and putatively deleted in chimpanzee. The primers successfully amplified the exons in humans and gorillas but yielded no products in chimpanzee, providing strong supporting evidence of the deletion.











