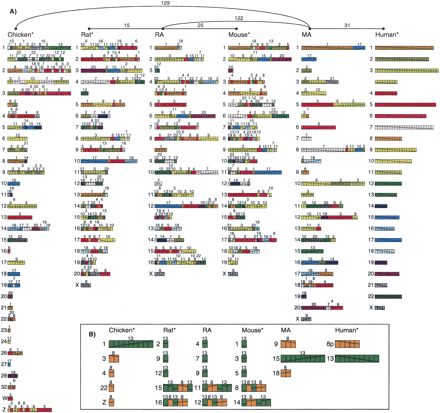
(A) Four pre-ancestors (marked with *), obtained after carrying out a maximal number of initial inversions, and two putative ancestors (murid rodent ancestor, RA, and mammalian ancestor, MA) for gene-based data in run gene7. Each genome is represented by an arrangement of 311 new synteny blocks obtained after carrying out the initial inversions and condensing the 586 initial four-way blocks. (B) Arrangement of the set of 14 blocks from HSA8p and HSA13, which are contiguous in the pre-ancestors of all species but are found on four different chromosomes in chicken, mouse, and rat. The two putative ancestral arrangements of these blocks, as recovered by MGR, are also shown.











