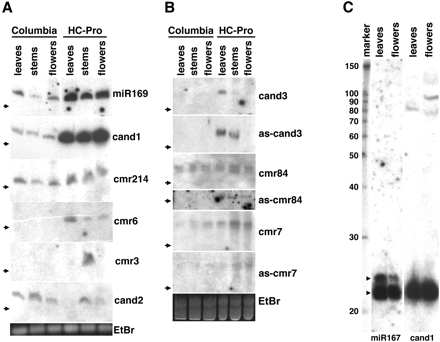
RNA gel-blot analysis of size-selected small RNAs from three tissues, leaves, stems, and flowers, from both wild-type Columbia plants and transgenic Columbia plants expressing the HC-Pro gene under the control of the 35S promoter. Signals were detected for 8 of the 13 selected miRNA candidates following hybridization of specific end-labeled DNA oligonucleotides. (Bottom) (EtBr) show representative RNA loadings for the set of replicate blots above.(A) Signals corresponding to ∼21 nt small RNAs. The candidates, cand1 and cmr214, show the expected increase in expression in the HC-Pro samples. Expression is detected only in HC-Pro lanes for cmr6 and cmr3, with higher expression in RNA isolated from leaf and stem tissue, respectively. For cand2, lower expression was observed in HC-Pro than the Columbia wild-type samples. Arrows indicate the position of a 23-nt DNA size marker that corresponds to RNAs of ∼20 nt. (B) StarFire probes were used to improve weaker signals corresponding to ∼24 nt small RNAs. Probes to the antisense sequences were also used to help identify whether the correct genomic strand was identified by findMiRNA (prefix `as-'). (C) Signals corresponding to end-labeled probes specific for miR167 and the cand1 candidate miRNA for RNA isolated from HC-Pro samples. An RNA size marker shown on the left edge has values shown in nucleotides. Larger bands that may represent precursor intermediates are evident for cand1. Two size classes of small RNA, ∼22 nt and ∼24 nt, are observed for the control, miR167 (arrowheads).











