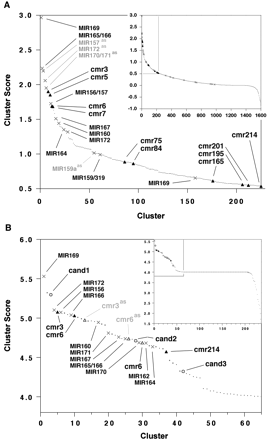
Ranked clusters of predicted candidate miRNA precursor sequences derived from two different findMiRNA experiments. Crosses (×) mark clusters containing previously reported miRNA precursors, filled triangles (▴) show clusters containing candidate miRNA precursors that were analyzed further, dots (·) indicate clusters of other candidate sequences. Clusters containing antisense sequences of previously reported miRNA precursors, for which the predicted target is erroneous, are shown in gray type. (A) Preliminary scan of 5701 transcripts. (Inset) All 1599 ranked clusters, with the magnified region delineated by gray borders. Clusters containing aligned sequences with 100% sequence identity across the entire alignment have a score of 0.0. (Magnified) Shows top-ranking clusters down to the cluster containing the cmr214 candidate precursor sequences. (B) Whole-genome scan, showing only ranked clusters that contain exact matches for the predicted miRNA sequence in rice. (Inset) All 236 ranked clusters, with the magnified region delineated by gray borders. (Magnified) Open circles (○) show candidates additional to those shown for preliminary scan. Small crosses (×) represent clusters containing some sequences that are also present in superior clusters (×) of previously reported miRNA precursors. Clusters containing aligned sequences with 100% sequence identity across the entire alignment have a score of 4.0.











