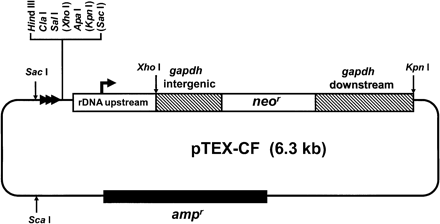
The chromosome fragmentation vector pTEX-CF. The vector was constructed by modifying the expression construct pRIBOTEX, as described in Methods. The gapdh-flanking sequences (hatched boxes) ensure correct processing of the neor transcript. Targeting fragments can be cloned in both orientations into the multiple cloning site (MCS). Bracketed restriction sites in the MCS are not unique. The location and direction of the repeated telomeric hexamers are indicated by horizontal arrowheads. The plasmid DNA (thin line) is derived from pBluescript and contains an ampr gene (filled box). Following insertion of the targeting fragment into the MCS, the construct can be used for transfection after linearization with the appropriate restriction enzyme.











