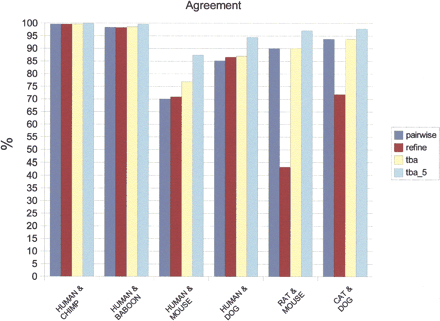
Agreement score of pairwise and multiple alignments produced by different aligners on a set of nine simulated mammalian sequences of length ∼50 kb. Pairwise results from BLASTZ were postprocessed to remove overlapped regions. Multiple aligners including refine and TBA use the same pairwise alignments. TBA_5 refers to alignments from TBA, but the agreement score allows mismatches within five base positions. Agreement scores of multiple aligners are measured from the pairwise alignments induced by pairs of species. All values are averaged over 50 sets of simulation sequences. Parameters used in the simulation and alignment programs are described in the text.











