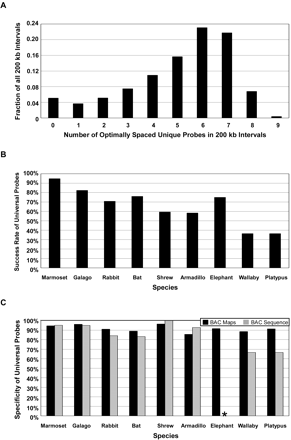
Characterization of the mammalian whole-genome universal hybridization probe set. (A) To estimate the fraction of the human genome associated with one or more mammalian universal probes, the number of unique probes selected using an optimal spacing parameter of 30 kb in all 200-kb windows (50-kb slide) across the genome, excluding intervals that contained a sequence gap in the human assembly, was determined. The result of that analysis was plotted as the fraction of all 200-kb intervals that included N number of optimally spaced unique probes. (B) To estimate the ability of universal probes to identify a BAC clone from mammalian libraries, a sample of n = 96 universal probes were hybridized to a panel of nine representative mammalian BAC libraries. The success rate, as defined by the percentage of universal probes that detected at least one positive BAC clone, for each of the nine species is indicated. (C) Specificity of the universal probes, as measured by physical mapping, and sequence analysis of individual isolated BAC clones is indicated. The number of BAC sequences included in the analysis for each species is as follows: marmoset, n = 21; galago, n = 20; rabbit, n = 19; bat n = 12; shrew, n = 16; armadillo n = 14; wallaby, n = 12; and platypus, n = 9. In the case of marmoset, shrew, and armadillo, probe specificity as measured by analysis of sequenced BAC clones was slightly higher than that of the probes themselves. Since multiple linked probes were included in the test set of probes, in these species many clones were positive for more than one probe and therefore facilitated a more accurate selection of orthologous versus nonorthologous clones for sequencing. *No BAC sequence is available for elephant.











