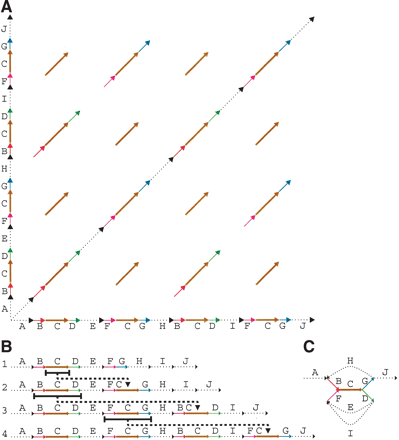
(A) Genomic dot-plot of an imaginary sequence with repeats containing sub-repeats. How many repeats are really present? The coloring shows the repeats from which it was constructed. In a real sequence, we would not know the sub-repeat structure and thus we would not be able to color the dot-plot. Also, because the repeats would not be perfect, there would be gaps, substitutions, and indels distorting each diagonal. In certain applications, there would be reverse diagonals, corresponding to alignments with opposite strands. (B) An imaginary evolutionary process leading from a repeat-free genome to the genome in A with four repeat copies. Each step duplicates a region and inserts it elsewhere. (C) Gluing repeated regions leads to the repeat graph of the final genome. Deleting the multiplicity 1 edges (shown dotted) in this graph leaves a single component, called a tangle or repeat. It consists of five edges B, C, D, F, G (shown solid) called sub-repeats. Every new duplication in B creates a more and more complicated tangle describing an evolving repeat structure. The graph structure of this tangle documents the evolutionary history of duplications.











