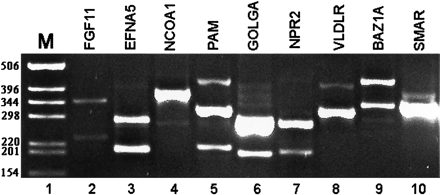
Experimental validation for the existence of alternative splicing in selected predicted exons. RT–PCR for 15 exons (detailed in Table 2), for which no EST/cDNA indicating alternative splicing was found, was conducted over 14 different tissue types and cell lines (see Methods). Detected splice variants were confirmed by sequencing. For nine of these exons a splice isoform was detected in at least one of the tissues tested. Only a single tissue is shown here for each of these nine exons. Lane 1, DNA size marker. Lane 2, exon 2 skipping in FGF11 in ovary tissue (the 344-nt and 233-nt products are exon inclusion and skipping, respectively). Lane 3, exon 4 skipping in EFNA5 gene in ovary tissue (exon inclusion 287 nt; skipping 199nt). Lane 4, exon 8 skipping in NCOA1 gene in placenta tissue (exon inclusion 377 nt; skipping 275 nt). Lane 5, exon 22 skipping in PAM gene in cervix tissue (exon inclusion 323 nt; skipping 215 nt). Additional upper band contains a novel exon in PAM. Lane 6, exon 9 skipping in GOLGA4 gene in uterus tissue (exon inclusion 288 nt; skipping 213 nt). Lane 7, exon 9 skipping of NPR2 gene in placenta tissue (282nt inclusion; 207nt skipping). Lane 8, intron 8 retention in VLDLR gene in ovary tissue (wild type 324 nt; intron retention 427 nt). Lane 9, alternative acceptor site in exon 12 of BAZ1A in ovary tissue (wild type 351 nt; alternative acceptor variant 265 nt). The uppermost band represents a new exon in BAZ1A, inserted between exons 12 and 13. Lane 10, alternative acceptor site in exon 7 of SMARCD1 in uterus tissue (wild type 353 nt; exon 7 extension 397 nt).











