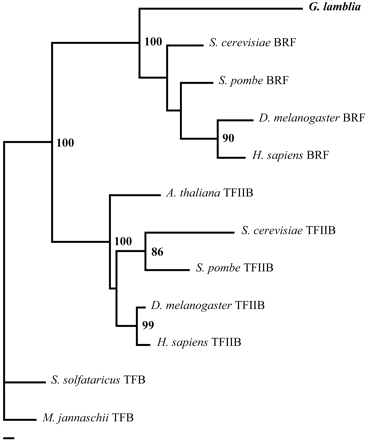
Phylogeny of archaeal TFB, eukaryal TFIIB, and eukaryal BRF transcription factors. Archaeal TFB sequences are used as the outgroup. Support values for the class V consensus tree (see Methods) are indicated next to each branch present in >50% of the significant topologies. Branch lengths of the consensus tree were estimated using PAML. The scale bar represents 10 amino acid replacements per 100 positions. Abbreviated sequence identifiers are Drosophila melanogaster, Giardia lamblia, Homo sapiens, Methanococcus jannaschii, Saccharomyces cerevisiae, Schizosaccharomyces pombe, and Sulfolobus solfataricus.











