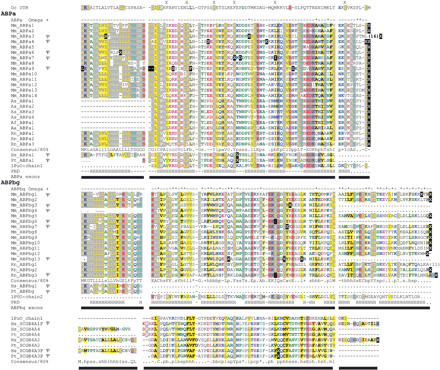
Amino acid sequence multiple alignment of rodent and primate ABPα and ABPβγ homologs, and primate secretoglobin homologs. Conceptual translations of both genes and pseudogenes (denoted by Ψ) are shown, with stop codons replaced by “X” symbols in white on black; codons containing a frame shift are also shown in white on black. Pseudo-exon sequences that could not be identified in genomic sequence are replaced by “.” characters; “-” represents a gap position. Multiple sequence alignments were produced by CLUSTAL W, manually adjusted, and colored using Chroma (Goodstadt and Ponting 2001) and an 80% consensus; gap positions were ignored in the calculation of a consensus sequence. Exons are shown as horizontal bars, and intron and exon boundaries were determined from alignments of the gene predictions and genomic DNA sequence using the UCSC genome browser. Protein secondary structure predicted by PHDSec (Rost and Sander 1993; PHD) shows four α-helices (H) in α-like chains, as has been demonstrated for cat Fel dI, rabbit uteroglobin, and human uteroglobin, and five α-helices in βγ-like chains. Codons predicted to be subject to positive selection by Codeml with a posterior probability of p > 0.9 in one model, and of at least p > 0.5 in one other model, are termed ω+ sites and are marked with an asterisk above the alignment. Positions of rabbit uteroglobin (UTG) residues predicted to interact with a bound ligand are each indicated by “X” above the aligned rabbit UTG sequence. (Mm) Mus musculus; (Hs) Homo sapiens; (Rn) Rattus norvegicus; (Pt) Pan troglodytes; (As) Apodemus sylvaticus; (Af) Apodemus flavicollis; (Aa) Apodemus agarius; and (Oc) Oryctolagus cuniculus.











