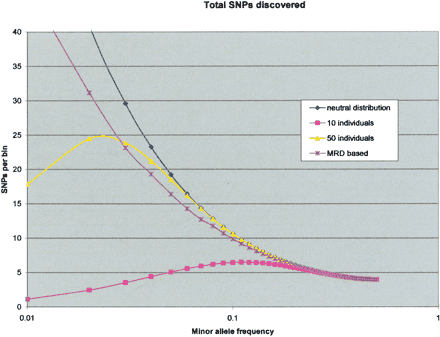
Sensitivity of MRD-based and traditional SNP discovery. We have modeled the proportion of SNPs that would be detected by various approaches. The y-axis is a relative measure of the number of SNPs, and the x-axis is the logarithm of the minor allele frequency. The model assumes a variation distribution consistent with neutral theory, and the total number of SNPs is shown in the blue curve. A perfect SNP discovery approach would have essentially the same curve. This neutral theory assumption is an approximation for at least three reasons. First, SNPs aggregate close together, more than expected randomly (Reich et al. 2002). Second, this aggregation is often generally due to variants on the same chromosome (Reich et al. 2002). Third, the variation distribution in exons is shifted toward the rarer alleles (Cargill et al. 1999; Halushka et al. 1999). The aggregation of SNPs would make the problem of two SNPs in the same amplicon more pronounced. This is balanced by the presence of LD and the occurrence of variants on the same chromosome, allowing for the enrichment of both SNPs together. The shift toward rare allele frequencies in the exons also should decrease the effect of the two SNPs in the same amplicon. Performance of traditional sequencing assumes perfect technical detection but takes into account the sampling error described above and requires two observations of the allele. We show the performance for sequencing 10 and 50 people (in pink and yellow, respectively). For the MRD-based SNP discovery (purple curve), we have taken into account the sensitivity determined from the BRCA experiment (Fig. 4). We also assumed that we would design our amplicons in such a way as to avoid validated SNPs in the public databases (to ameliorate the two SNPs in one amplicon effect). The proportion of SNP detected in the model has taken into account the expected times two SNPs are present on the same amplicon after avoiding database-validated SNPs in the amplicon design phase.











