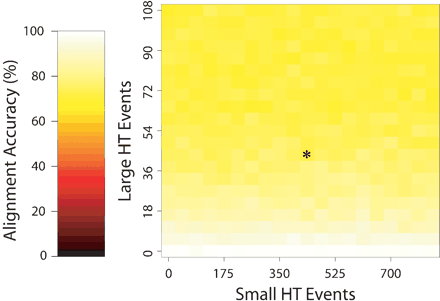
The performance of Mauve when aligning sequences evolved with rates similar to those observed among the group of nine enterobacteria. In this experiment, the substitution, indel, and inversion frequencies were held constant at rates similar to those observed in the enterobacteria. The asterisk (*) denotes the combination of large and small horizontal transfer rates observed in the enterobacteria. As the rate of large horizontal transfer increases, the amount of lineage-specific sequence relative to backbone grows. Because Mauve cannot align large lineage-specific regions, the alignment score drops. When scored only on regions considered backbone sequence, the accuracy is consistently above 98%.











