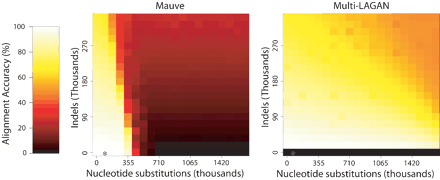
The performance of Mauve (left) and Multi-LAGAN (right) when aligning sequences evolved with increasing amounts of nucleotide substitution and indels. The multi-MUM anchoring technique used by Mauve limits its ability to align distantly related sequences. Multi-LAGAN version 1.2 did not complete the alignments of genomes without indels, resulting in the black row at the bottom. The substitution and indel rate observed in the enterobacteria is denoted by an asterisk (*).











