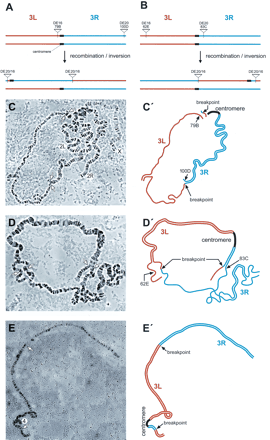
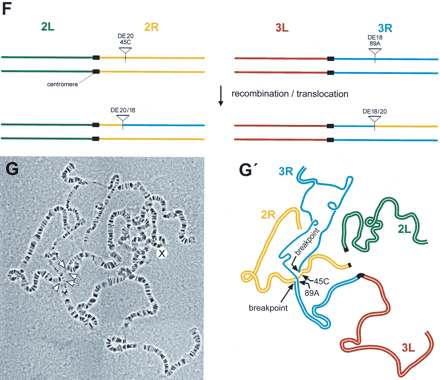
Cytology of chromosome 3 inversions (A–E) and chromosome 2; three translocations (F, G). Scheme of the generation of inversions In(3LR)DE16d*-79A;DE20z -100D derived from stock α (A) and of In(3LR) DE20h-62E; DE16c-83C derived from stock β (B). (C, C′) Polytene chromosomes and schematic representation from In(3LR)DE16d-79A;DE20z -100D/+ heterozygotes. (D, D′) Polytene chromosomes and schematic representation from In(3LR) DE20h-62E; DE16c-83C/+ heterozygotes, and (E, E′) from In(3LR) DE20h-62E; DE16c-83C homozygotes. Note the unequal length of chromosomal arms of the inverted chromosomes. (F) Scheme of the generation of chromosome 2; three translocations. (G, G′) Polytene chromosomes and schematic representation from T(2; 3) DE18c-45C; DE20d-89A/+ heterozygotes. Lowercase letters arbitrarily denote the isolated P-element insertion.











