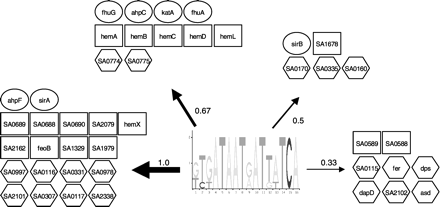
Schematic representation of the predicted fur regulog. The logo represents the pattern that was obtained by phylogenetic footprinting. The arrows that connect the pattern and the groups of genes under the control of the pattern indicate the relative conservation score of the gene, the thickest arrows belonging to the most-conserved genes, for which we thus have a high confidence that they belong to the regulog. Ovals indicate known members in S. aureus. Rectangles indicate genes that may be suspected to be in the Fur regulog, on the basis of their sequence similarity to proteins in other organisms that have been shown to be regulated by Fur. Hexagonal boxes represent new members of the regulog. This means that they are predicted by Regulogger to be regulated by Fur, but no experimental evidence exists.











