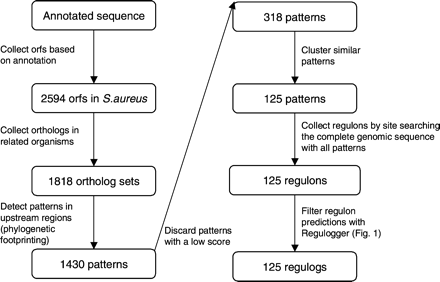
Schematic representation of the strategy to identify regulogs in S. aureus. From the genomic sequence, protein-coding regions are identified. For all proteins, orthologs in other genomes are defined. These ortholog sets are used for phylogenetic footprinting, in which Gibbs sampling is run on upstream regions of sets of orthologous genes to obtain putative regulatory motifs (e.g., binding sites). Low-scoring patterns are filtered and patterns with similar sequences are clustered. For each pattern, the putative regulon in S. aureus is defined. These predicted regulons are filtered with the Regulogger method described in Figure 1. This produces a set of regulogs, conserved regulons, in S. aureus.











