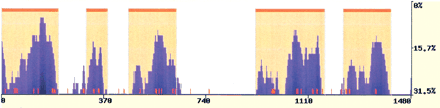
Figure 5
eShadow application in multiple protein alignment analysis. Human, baboon, cow, sheep, mouse, rat, and rabbit CFTR proteins were aligned using eShadow, and highly homologous regions were predicted using the HMMI module (0.77/0.65/0.1). Mutations known to cause cystic fibrosis (http://www.genet.sickkids.on.ca/cftr) are mapped to the CFTR MPA and annotated as red tick marks.











