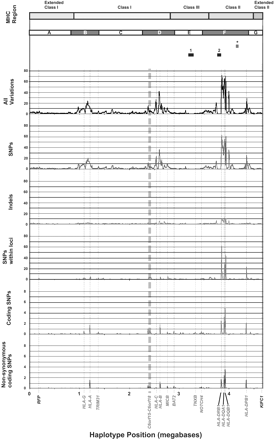
Positional distribution of variations between COX and PGF MHC sequences. MHC sequences were divided into 10-kb bins, and variations were calculated in each bin. Results are expressed as variations per 1 kb. A locus is defined as all genomic DNA between the 5′-start of a gene to the 3′-end. Boundaries of the class I, II, and III regions are shown. The positions of genes RFP and KIFC1 that define the ends of the MHC haplotype sequencing project are indicated in black. Other genes with five or more SNPs between the haplotypes are labeled in blue. HLA-C is also labeled. Regions of interest are labeled above the graph. Regions 1 and 2 are the RCCX module and the HLA-DRB region, respectively. (*) The TAP to HLA-DMB region, which shows little variation between haplotypes. Blue bars B, D, and F label regions in which variation is thought to be classical MHC class I and class II gene-associated.











