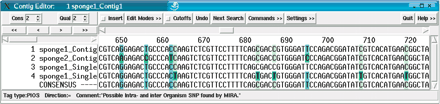
The last (optional) step of the EST assembly consists of the input sequences being given strain information to show the effect when two different organism strains (named sponge1 and sponge2) are sequenced and analyzed. In this example, miraEST classified the SNPs into two categories: PROS (shown in light blue) for SNPs that occur only between strains/organisms (e.g., column 661) and PIOS (shown in light green) for SNPs that occur both within a strain as between different strains (e.g., column 662). Interestingly enough, most of the SNPs shown in this example will not cause a change in the amino acids of the resulting protein, with one notable exception, the SNP of sponge1_singlet4 at base position 662 causes a TAA codon to be expressed, which is a stop codon. The SNPs of the same sequence at position 686 and 707 would cause mutations in the amino acid sequence, but are, because of the TAA mutation earlier, in the3′ UTR of this particular mRNA transcript.











