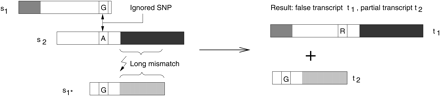
Example of a misassembled transcript when SNPs are disregarded. Assembly of three input sequences are shown at left; the resulting transcripts of this assembly are shown at right. The three sequences s1, s1*, and s2 contain different homologous parts, represented by the different shades of gray, and exactly one SNP position. A normal assembly algorithm will assemble first s1, then s2 (because of the long overlapping alignment in the white part), and then might try to align s1*, but fail because of the large mismatch. The SNP position with G in sequence s1 and A in s2 is treated as typical noise in the alignment algorithms and ignored. The resulting transcript sequences are therefore wrong, as they do not represent the sequences found in vivo: t1 is a mix of two transcripts and does not code a true protein.











