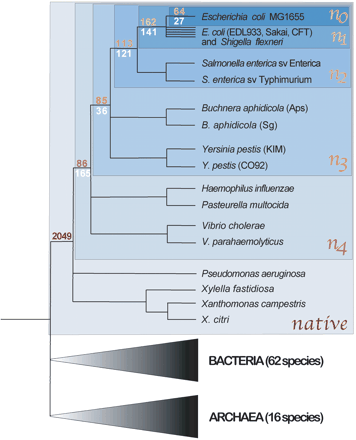
Distribution of clade-specific genes at different phylogenetic depths within the γ-Proteobacteria. The topology of the tree is based on Lerat et al. (2003), and successive blue boxes (n0–n4, native) encompass the clades considered in the present study. Numbers of ORFans (yellow/red) and HOPs (white) in the E. coli MG1655 genome specific to n0–n4 are shown at the basal nodes of each clade. The number of native genes (n = 2049) corresponds to genes in the E. coli MG1655 genome that are present in at least one member of each clade. Species numbers of Bacteria and Archaea denote all those included in BLASTP searches.











