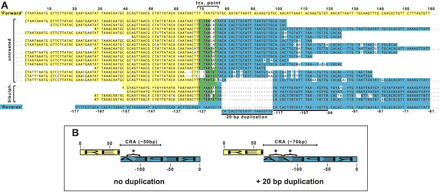
The structure of minichromosomal 177-bp repeat inversions. (A) Sequence of 16 nonredundant repeat inversions isolated from untreated genomic DNA and four nonredundant inversions from bisulfite-treated DNA. Because bisulfite treatment introduces genetic code degeneracy, these sequences consist of residues {A, G, Y} for “A” strand amplicons or {C, T, R} for “B” strand amplicons. Sequences are aligned against “forward” (yellow) or “reverse” (blue) orientations of the 177-bp repeat. Around the inversion point are nucleotides that match both orientations (green). A 20-bp subrepeat duplication found in some inversion point sequences is marked with a solid line. (B) Schematic representations of the isolated repeat inversions highlighting the central region of asymmetry (CRA). This CRA is ∼50 bp or ∼70 bp depending on duplication of the 20-bp subrepeat (starred).











