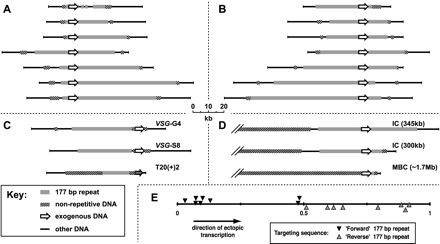
(A–D) Physical maps of 20 T. brucei chromosomes deduced with the aid of ectopic plasmid insertions. Here 17 complete minichromosomes were mapped from integration events targeted to (A) “forward” 177-bp repeat; (B) “reverse” 177-bp repeat; and (C) VSG-G4, VSG-S8, and an orphan rDNA promoter (Zomerdijk et al. 1992). Partial maps were also produced from integration events into 177-bp repeats occurring on two ICs and one MBC (D). The IC of 345 kb was targeted using “forward” repeats, whereas the IC of 300 kb and MBC of ∼1.7 Mb were targeted using “reverse” repeats. Partial maps only cover DNA up to ∼100 kb from the site of integration. All chromosomes are arranged such that vector transcription runs left to right. The resolution of the maps decreases with increasing distance from the integrated vector (see Fig. 2C). Combining data from several gels, we estimate the standard error in measurements of size around the site of insertion to be <20 bp, whereas that 50 kb–100 kb away is ∼2 kb. (E) Position of the integration events along the 177-bp core when targeting “forward” or “reverse” 177-bp repeats. Positions are normalized with respect to the total length of repetitive core region (excluding vector sequence) for each individual chromosome.











