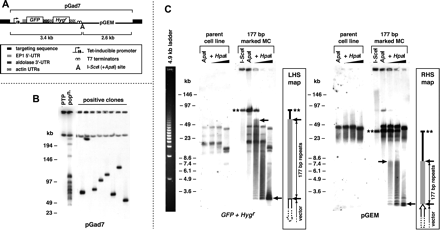
(A) Anatomy of vectors used in this study to ectopically mark T. brucei chromosomes. Vectors were derived from pGad7 and differ only in targeting sequence: “forward” 177-bp repeat; “reverse” 177-bp repeat; VSG-G4 or VSG-S8. The recognition site for the extremely rare-cutting endonuclease I-SceI is shown. A unique ApaI site exists immediately adjacent to this site. (B) Integration into T. brucei small chromosomes by targeting the 177-bp repeat. The distribution of vector (pGad7) integration observed in a population of uncloned positive transformants (popn.) closely matches the distribution of the 177-bp repeat (see Fig. 1) and is the result of single integration events in many cells, as seen when the population is cloned. The pGad7 probe also hybridizes to pGEM-derived sequence on an MBC in the parental PTP cells. (C) Mapping 177-bp repeats regions on ectopically marked MCs. Genomic DNA from a clone marked at a 177-bp repeat was digested to completion with either I-SceI or ApaI endonucleases, both of which cut uniquely within the exogenous plasmid sequence. DNA digested with ApaI was then digested with 0, 0.08, 0.4, or 2 units of HpaI. Hybridization of PFGE-separated DNAs to either GFP and Hygr probes or pGEM probe is shown. Double stars indicate DNA uncut by HpaI. Arrows denote the extent of the 177-bp repeat regions. Note that there is cross-hybridization of the probes to sequences in the parental PTP cell line, particularly pGEM, which is present in PTP cells. The migration profile of a 4.9-kb ladder (left panel) under the same conditions gives an indication of gel resolution.











