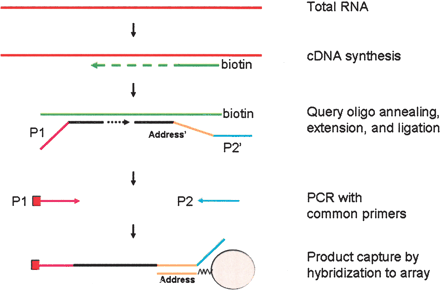
DASL: a cDNA-based assay for RNA profiling. RNA is converted to cDNA, using biotinylated oligo-d(T)18 and random hexamers, and immobilized to a streptavidin-coated solid support. Two oligonucleotides are designed to interrogate each target site on the cDNA. The upstream oligonucleotide consists of two parts, the gene-specific sequence and a universal PCR primer sequence (P1) at the 5′-end. The downstream oligonucleotide consists of three parts, the gene-specific sequence, a unique address sequence which is complementary to a capture sequence immobilized on the array, and a universal PCR primer sequence (P2′) at the 3′-end. A single address sequence is uniquely associated with a single target site. The upstream oligonucleotide hybridizes to the targeted cDNA site, extends and ligates to its corresponding downstream oligonucleotide to create a PCR template that can be amplified with universal PCR primers (P1 and P2). The PCR products, which are fluorescently labeled by incorporation of the 5′-labeled primer P1, are hybridized to capture sequences on the beads in the array, and fluorescence intensity is measured for each bead.











