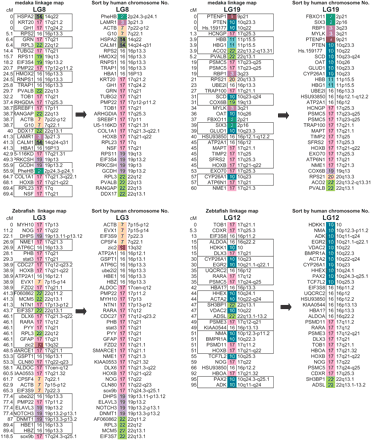
The medaka linkage map of LG8 and LG19 and the orthologous zebrafish chromosomes, LG3 and LG12. Both pairs have HoxB clusters, and the orthologs are sorted according to human chromosome order. The linkage data for zebrafish was obtained from Heat Shock Diploid Cross, Maps and Data, in the Stanford Zebrafish Genome Project (http://zebrafish.stanford.edu/genome/Frontpage.html). The locations of 23 human chromosomes are distinguished by 23 colors, and the location of human orthologous genes are indicated by color and human chromosome numbers. Markers in boxes showed no recombination. Gene order was scrambled by apparent inversions occurring since the medaka and human divergence. After sorting (canceling the effects of inversions), similar domain structures are observed both in LG8 and LG19, indicating that these chromosomes arose from the duplication of a single ancestral chromosome. The markers which have two possible locations in the same LG are not used in this figure but are used in Figure 4.











