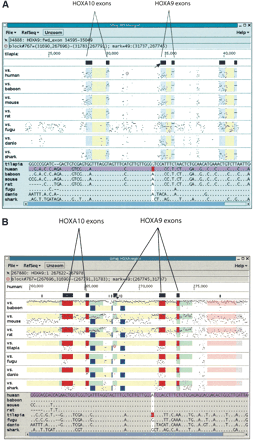
A threaded blockset for vertebrate HoxA regions, displayed in our interactive blockset viewer Gmaj. (A) The red circle marks a position of interest where the tilapia reference sequence aligns with human. The block containing this position is highlighted in red in all of the alignment panels. Color underlays are blue for exons in the reference sequence and yellow for introns, and the exons are also represented as icons above the alignments. At the top of the Gmaj window, two status lines describe the positions of the mouse pointer and the red circle, respectively. Individual nucleotides for the selected block are displayed in the bottom pane, with the marked position highlighted. (B) The same region projected onto the human sequence. The underlays for human include (green) for EST evidence, (dark blue) for antisense RNA, and (red) for coding sequences. The conserved element from A is part of an alternative 5′-end identified by homology to a human EST from TIGR.











